Mathematical Modeling Approaches to Understanding Severe Acute Respiratory Syn-drome Coronavirus 2 (SARSCoV-2) DNA Sequences Linked Coronavirus Disease (COVID-19) for Discovery of Potential New Drugs
ABSTRACT
The novel coronavirus called SARS Coronavirus-2 is associated with severe acute respiratory syndrome. At present, in-tensive care unit treatment and specialized treatment have been not available [1]. Mathematical modeling approaches to understanding the evolution of the DNA sequences of SARS0CoV-2, is an efficient way to design biochemical components interacting with the virus DNA structure. This study aimed to identify specific sets of nucleotides partially “mirror repeats” sets of nucleotides in DNA sequences of SARS0CoV-2. It is suggested that understanding of the “mirror repeats” (Generalized Smarandache Palindrome Sequence (S. Palindrome)) (Dawoudi 2018) sets of nucleotides can be informative in revealing virus gene functions. Under-standing more about coronavirus genome functions and transcriptomic level might allow drug designer to design exact treatments
KEYWORDS
COVID-19; SARS0CoV-2; S. Palindrome; Motif; Oligonucleotide, Frequency distribution; Adaptation studies
INTRODUCTION
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) was named by the International Committee on Taxonomy of Viruses (ICTV) [2] is causes of new pandemic disease called coronavirus disease of 2019 (COVID-19) [3]. The virus is contributed to respiratory system and ‘acute lung symptoms’ Jiabao et al. [4]. The fatality rate of COVID-19 has been correlated with hypertension, diabetes, coronary heart disease, cerebral infarction, and chronic bronchitis Sheng [5] Up to now “there is no specific treatment available’ Phadke [6]. Nevertheless, ‘the current evidence of potential therapeutic agents, such as Barlow et al. [7] Chloroquine and Hydroxychloroquine/Plaquenil [8], Favilavir [9], Iopinavir plus Ritonavir [10] Remdesivir, Emetine, Homoharringtonine [11], Tocilizumab [12], Peripheral lymphocyte [13] and Angiotensin Receptor Blockers (ARBs) [14] Interferon, Ribavirin, Tocilizumab and Sarilumab [15] are promising in blocking infection. However, according to [7]. ‘There is currently no widely accepted standard of care in the pharmacological management of patients with COVID-19 [16]. To obtain efficient treatment, analysis of DNA sequences of SARS0CoV-2 for development and discovery of potential new clinical treatment by mathematical modeling methods, are the essential ways for designing of new drugs. This study has been focused on ‘mirror repetitive elements, in DNA sequence of SARS0CoV-2. ‘The sequence of nucleotides which are call Generalized Smarandache Palindrome Sequence (GSPs) or mirror repeats is defined as follows: nucleotides of the form r1 r2 r3 … rn-1 rn rn-1 … r3 r2 r1 with n > 0, where r1, r2, r3, … , rn are consisting various nucleotide of A, C, G or T [17]. According to Paulson 2018 many disease ‘are caused by expansions of simple sequence repeats dispersed throughout the human genome [18]. This study has demonstrated the range distribution of ‘mirror repetitive elements’ in SARS0CoV-2 sequence and has shown the possible relevant causes of COVID-19.
METHODS AND MATERIALS
To analyses of complete genome of SARS0CoV-2, our genomic data has been provided from National Center for Biotechnology. Genomic analyses of SARS0CoV-2 were per-formed by PALINDROM_ FINDER tools. This program has developed for detecting and analysis of possible mirror repetition in gene sequence. This gene come from FL, USA and was submitted on 16-FEB-2020 in the Public Health and Infection Research Group, with ID in GenBank: MT072688.1. This research study was investigated the frequency of “mirror repeats” of nucleotide (Generalized Smarandache Palindrome Sequence (S. palindrome)). Dawoudi in SARS0CoV-2 sequence about values of relative frequency (percentage of observation). In this research correlation between SARS0CoV-2 gene and these peptides have been considered. There are seven steps to find S. palindrome in SARS0CoV-2 (Figure 1).
Step 1: Identification of Dinucleotide S. palindrome
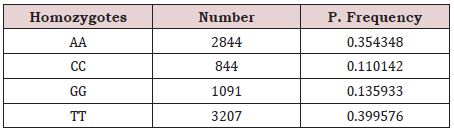
In this step four Homozygotes (‘The two alleles are in the same state’) [19] including 2844 AA, 884 CC, 1091 GG and 3207 TT was recognized. (Table 1) Maximum frequency is 0.399576 (TT) and minimum frequency 0.110142 (CC).
Step 2: Identification of Trinucleotide S.Palindrome
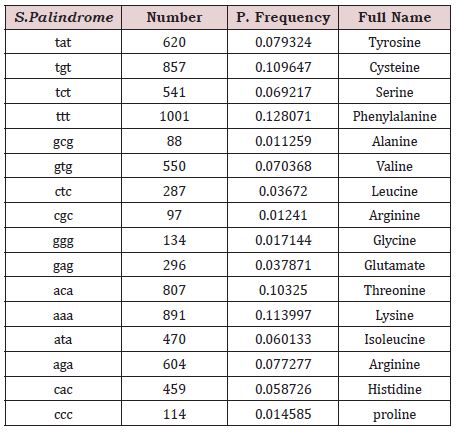
The total numbers of trinucleotide S.Palindrome are 7816, including: Tyrosine, Cysteine, Serine, Phenylalanine, Ala-nine, Valine, Leucine, Arginine, Glycine, Glutamate, Threo-nine, Lysine, Isoleucine, Arginine, Histidine and Proline. The maximum sampling frequency belong to Phenylalanine (TTT) rate 0.128071 and minimum sampling frequency rate is 0.01241 belong to Arginine (Table 2).
Step 3: Identification of Tetranucleotide S.Palindrome
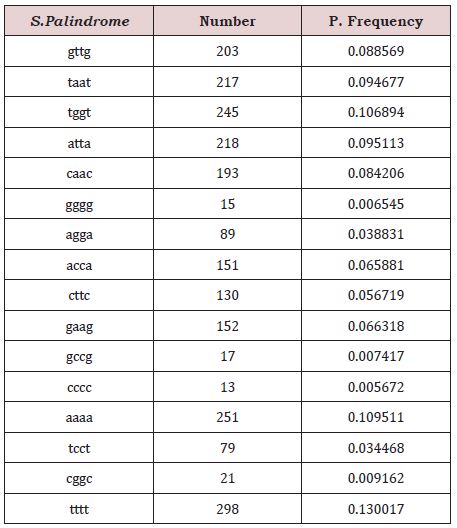
A fragment of gene with 4 base pairs mirror repetition (e.g. TAAT). The total number of Tetranucleotide S.Palindrome in SARS0CoV-2 were 2292. The maximum observed frequency was 298 TTTT and minimum observed frequencies is 13 CCCC (Table 3).
Step 4: Identification of Pentanucleotide S.Palindrome
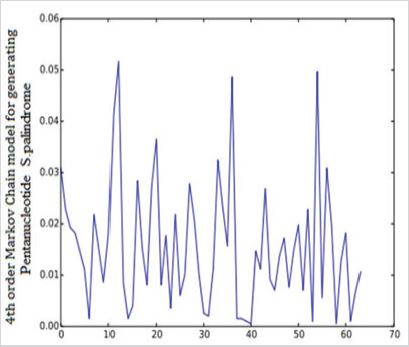
A fragment of gene with 5 base pairs mirror repetition (e.g. AGGGA). The total number of Pentanucleotide S.Palindrome in SARS0CoV-2 was 1973. The maximum observed frequen-cy was 102 TTGTT and minimum observed frequencies are 1 CGAGC and 1 GGGGG (Figure 2).
Step 5: Identification of Hexanucleotide S.Palindrome
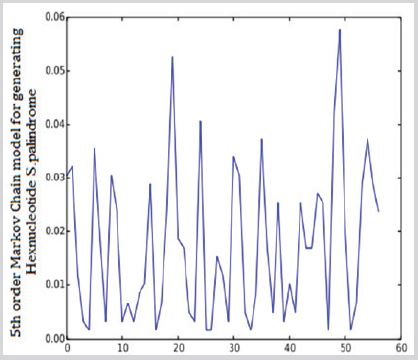
A fragment of gene with 6 base pairs mirror repetition (e.g. TGTTGT). The total number of Hexanucleotide S.Palindrome in SARS0CoV-2 was 590. The maximum observed frequency was 34 TGTTGT and minimum observed frequencies are 1 AGGGGA and 1 CTCCTC and 1GCCCCG and 1 TCCCCT and 1 TAGGAT and 1 AGCCGA and 1 GATTAG (Figure 3).
Step 6: Identification of Heptanucleotide S.Palindrome
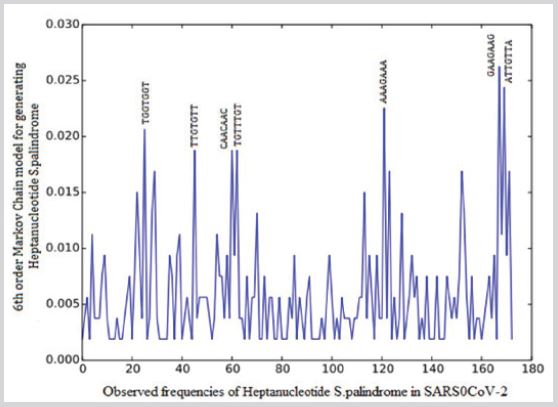
A fragment of gene with 7 base pairs mirror repetition (e.g. TGTTTGT). The total number of Heptanucleotide S.Palindrome in SARS0CoV-2 was 533. Maximum observed frequencies of Heptanucleotide S.Palindrome in SARS0CoV-2 is 14 AGGAGGA (Figure 4).
Step 7: Identification of Octa nucleotide S.Palindrome
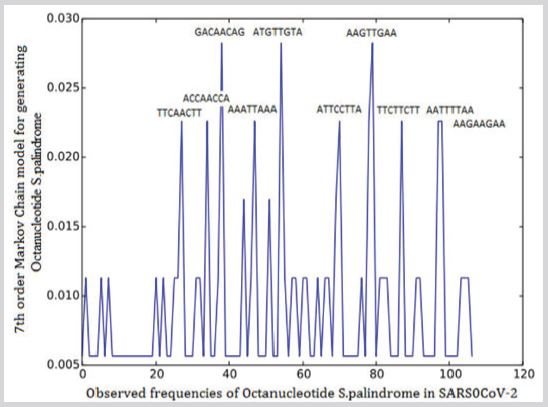
A fragment of gene with 8 base pairs mirror repetition (e.g.). The total number of octa nucleotide S.Palindrome in SARS0CoV-2 was 177. Maximum observed frequencies are AAGTTGAA, ATGTTGTA and GACAACAG (Figure 5).
RESULTS
The study has found 4 Dinucleotide in step 1, 16 Trinucleotide in step 2, 16 Tetranucleotide in step 3, 64 Pentanucleotide in step 4, 57 Hexnucleotide in step 5, 173 Heptanucleotide in step 6 and 107 Octanucleotide S.Palindrome in step 7. There are examples shows the crucial role of these peptides in human genes. In 2019 Szpiech and colleagues suggested that Runs of homozygosity (ROH) ‘are associated with an inflation of deleterious homozygous variation [20]. They have suggested that ‘African haplotype backgrounds may play a particularly important role in the genetic architecture of complex diseases [21]. One year later Wang et al. [12] have shown that ‘E1021K homozygous mutation in PIK3CD leads to activated PI3K-Delta Syndrome 1(immunodeficiency disease) [22]. In other study in 2017 Guo and et al claimed that ‘Trinucleotide repeat containing 6c (TNRC6c) is essential for microvascular maturation during distal airspace sacculation in the developing lung.’ [23]. The study was conducted by Chuang and et al in 2018 suggested ‘that the consensus SOX2 binding sequence, (T/A) TTGTT, could regulate the expression of COL1A1 [24]. Their finding ‘show evidence of BP as a potential therapeutic treatment in pulmonary fibro-sis [25]. ‘Zhang and colleagues found that the binding sites of circRNAs for some RBPs exhibit common patterns, such as the “GAAGAAG’” motif common among several RBPs including Argonaute 2(Ago2) (associated with Colorectal Cancer and Eunuchism [26] and α-ketoglutarate-dependent dioxygenase alkB homologue 5 (ALKBH5) (associated to Retinitis Pigmentosa 71 and Smith Magenis Syndrome.
As a result there are significant evidence that the higher rate observed frequencies of S.Palindrome appear in Step 6 and 7. The maximum observed frequencies of Heptanucleotide S.Palindrome GAAGAAG(14), ATTGTTA(13), AAAGAAA(12), TGTTTGT(11), CAACAAC(10), TGGTGGT(10). The top three significant Heptanucleotide S.Palindrome on SARSCoV-2 are: GAAGAAG, CAACAAC and TGGTGGT. Peptide GAAGAAG is included two consecutive GAA. In 2011 Tsuda and et al ‘revealed that the hTra2-β RRM strongly binds to the [5′-(GAAGAA)-3′] sequence. Peptide CAACAAC is included two consecutive CAA. identified GAAGAA ‘as the potential binding sites of SRSF10 within the alternatively spliced or flanking exons by using the SELEX approach [30,31]. Peptide TGGTGGT is included two consecutive TGG. In other experiment Rosani 2017 claimed that ‘Among the 104 invertebrate dsDNA viruses present in the dataset, only a truncated chi-motif (TGGTGG). Chuzhanova 2009 was widely enriched (in around 50% of the viruses, including OsHV-1 but not AbHV-1-AUS).
DISCUSSION
In this study, mathematical modeling approaches to understanding mechanism of SARSCoV-2 was introduced. This model has been bested on S. Palindrome algorithm. There are numerous mathematical models are developed as identification of genomic function. The following computational modeling of SARSCoV-2 sequence are proposed to acquiring better understanding of genomic function in COVID-19, e.g. ‘nth Order Markov Chain (nth- OMC) Mapping’ and ‘Biological Model of Distribution of Prime Numbers and Using Complex Network’.
CONCLUSION
Those characteristics of oligonucleotide and peptides that have been mentioned in this study are mostly related to Non-viral genes. However, exploring the interface between human gene function and coronaviruses gene structure, could be the effective way to better understanding of SARS0CoV-2 func-tions. This ‘adaptation studies’ [33] might be efficiency way for understanding aetiology of COVI-19 and ‘and provides potential new targets for designing drugs against’ [34] SARS0CoV-2 functions.
ACKNOWLEDGEMENT
I would like to thank my wife Afrouz Zibaei, Candida doctor at MMU for her inspiration and constant love, exhortation and support. Many thanks go to my Prof Richard Bayford, Professor of Biophysics & Engineering for providing me a unique chance to pursue my doctoral studies in Bioinformatics at Middlesex University London. The author would also like to thank Erika Andrea Kiss for her kindness, continuous motivation and encouragement.
REFERENCES
- Tarek Mohamed Abd El-Aziz, James DS (2020) Recent progress and challenges in drug development against COVID-19 coronavirus (SARSCoV- 2) an update on the status. Infect Genet Evol 83: 104327.
- Kit San Yuen, Zi WY, Sin YF, Chi PC, Dong YJ (2020) SARS-CoV-2 and COVID-19: The most important research questions. Cell Biosci 10: 40.
- Firas AR, Mazhar S Al Zoubi, Ghena AK, Dunia MS, Amjad D Al-Nasser (2020) SARS-CoV-2 and coronavirus disease 2019: What we know so far. Pathogens 9(3): 231.
- Jiabao Xu, Shizhe Z, Tieshan T, Abualgasim EA, Wan Z, et al. (2020) Systematic comparison of two animal-to-human transmitted human coronaviruses: SARS-CoV-2 and SARS-CoV. Viruses 12(2): 244.
- Sheng QD, Hong JP (2020) Characteristics of and public health responses to the corona virus disease 2019 outbreak in China. J Clin Med 9(2): 575.
- Phadke M, Saunik S (2020) COVID-19 treatment by repurposing drugs until the vaccine is in sight. Drug Dev Res.
- Barlow A, Landolf KM, Barlow B, Yeung SYA, Heavner JJ, et al. (2020) Review of Emerging Pharmacotherapy for the treatment of coronavirus disease 2019. Pharmacotherapy 40(5): 416-437.
- Duddu P (2020) Coronavirus treatment: Vaccines/drugs in the pipeline for COVID-19. Clinical Trials.
- Neil M (2020) Lopinavir-Ritonavir was not effective for COVID-19. Journal Watch.
- Choy KT, Wong AY, Kaewpreedee P, Sia SF, Chen D, et al. (2020) Remdesivir, lopinavir, emetine, and homoharringtonine inhibit SARSCoV- 2 replication in vitro. Antiviral Res 178: 104786.
- Michot JM, Albiges L, Chaput N, Saada V, Pom-meret F, et al. (2020) Tocilizumab, an anti-IL6 receptor antibody to treat Covid-19-related respiratory failure: a case report. Ann On-col.
- Wang F, Nie J, Wang H, Zhao Q, Xiong Y, et al. (2020) Characteristics of peripheral lymphocyte subset alteration in COVID-19 pneumonia. J Infect Dis 221(11): 1762-1769.
- Goldstein MR, Poland GA, Graeber CW (2020) Are certain drugs associated with enhanced mortality in COVID-19? QJM: An International Journal of Medicine.
- Jordan BR (2001) DNA Arrays for expression measurement: An historical perspective. In: Jordan BR (eds) DNA microarrays: gene expression applications. Principles and practice. Springer, Berlin, Heidelberg.
- The correlations between C9ORF72-linked depressive pseudodementia and apolipoprotein E (ApoE)-linked alzheimer disease (AD) using generalized smarandache palindrome sequence mapping Algorithm.
- Paulson H (2018) Repeat expansion diseases. Handb Clin Neurol 147: 105-123.
- Szpiech ZA, Mak ACY, White MJ, Hu D, Eng C, et al. (2019) Ancestrydependent enrichment of deleterious homozygotes in runs of homozygosity. Am J Hum Genet 105(4): 747-762.
- Wang Y, Chen X, Yang Q, Tang W, Jia Y, et al. (2020) E1021K Homozygous Mutation in PIK3CD Leads to Activated PI3K-Delta Syndrome 1. J Clin Immunol 40(2): 378-387.
- Guo H, Kazadaeva Y, Ortega FE, Manjunath N, Desai TJ (2017) Trinucleotide repeat containing 6c (TNRC6c) is essential for microvascular maturation during distal air space sacculation in the developing lung. Dev Biol 430(1): 214-223.
- Chuang HM, Ho LI, Huang MH (2018) Non-canonical regulation of type I collagen through promoter binding of sox2 and its contribution to ameliorating pulmonary fibrosis by Butylidenephthalide. International Journal of Molecular Sciences. 19(10): 3024.
- Gene cards.
- Huang A, Zheng H, Wu Z, Chen M, Huang Y (2020) Circular RNA-protein interactions: functions, mechanisms and identification. Theranostics 10(8): 3503-3517.
- Kengo T, Tatsuhiko S, Kanako Kk, Mari T, Fahu H, et al. (2011) Structural basis for the dual RNA-recognition modes of human Tra2-β RRM. Nucleic Acids Res 39(4): 1538-1553.
- Lin JC (2015) Impacts of alternative splicing events on the differentiation of adipocytes. International Journal of Molecular Sciences16(9): 22169- 22189.
- Feng Y, Valley MT, Lazar J, Yang AL, Bronson RT, et al. (2009) SRp38 regu-lates alternative splicing and is required for Ca2+ handling in the embryonic heart. Dev. Cell 16(4): 528-538.
- Umberto R, Paola V (2017) Oyster RNA-seq data support the development of malacoherpesviridae Genomics 8:1515.
Article Type
Research Article
Publication history
Received date: May 07, 2020
Published date: May 27, 2020
Address for correspondence
Mohammad Reza Dawoudi, Prospective doctoral student at Middlesex University, London
Copyright
©2020 Open Access Journal of Biomedical Science, All rights reserved. No part of this content may be reproduced or transmitted in any form or by any means as per the standard guidelines of fair use. Open Access Journal of Biomedical Science is licensed under a Creative Commons Attribution 4.0 International License
How to cite this article
Mohammad Reza D. Racism: Mathematical Modeling Approaches to Understanding Severe Acute Respiratory Syn-drome Coronavirus 2 (SARSCoV-2) DNA Sequences Linked Coronavirus Disease (COVID-19) for Discovery of Potential New Drugs. 2020 - 2(3) OAJBS.ID.000173.
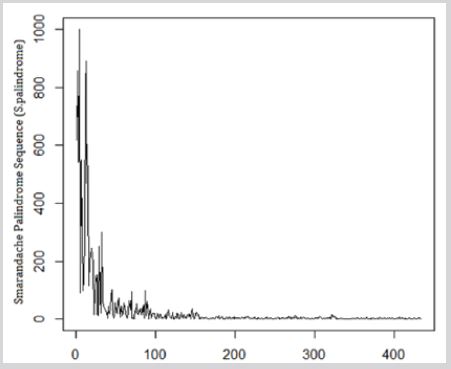
Figure 1: Observed frequencies of 3,4,5,6,7 and 8 S.Palindrome in SARS0CoV-2.
Figure 2: Observed frequencies of pentanucleotide S.Palindrome in SARS0CoV-2.
Figure 3: Observed frequencies of Hexanucleotide S.Palindrome in SARS0CoV-2.
Figure 4: Observed frequencies of Heptanucleotide S.Palindrome in SARS0CoV-2.
Figure 5: Observed frequencies of Octanucleotide S.Palindrome in SARS0CoV-2.
Table 1: Observed frequencies of Homozygotes in SARCOCoV-2.
Table 2: Observed frequencies of Trinucleotide S.Palindrome in SARCOCoV-2..
Table 3: Observed frequencies of tetranucleotide S.Palindrome in SARCOCoV-2.